
November 07, 2022 | 5 minutes read
Pandas is an amazing library for the analysis of low-dimensional labelled data — if the data you’re dealing with can be categorized as “rows and columns”, Pandas has you covered.
Most Geospatial data however does not fit into the row-column paradigm. It is often multidimensional in nature.
NetCDF (Network Common Data Form) is a set of software libraries and self-describing, machine-independent data formats that support the creation, access, and sharing of array-oriented scientific data.
Let’s go through some NetCDF vocabulary first-
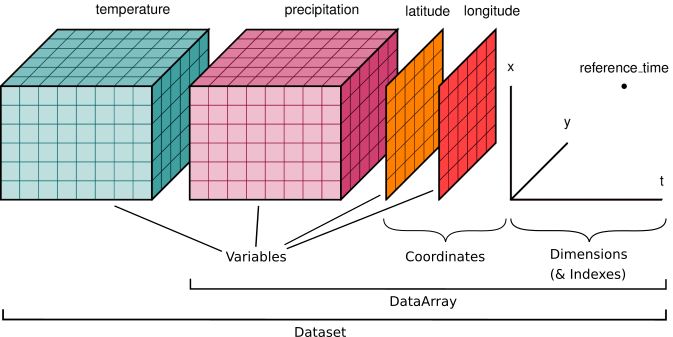
- Dimensions: these are used to specify the shape of one or more of the multi-dimensional variables contained in a NetCDF file. You can think of them as physical quantities. For eg. latitude, longitude, depth, time, altitude etc.
- Data Variables: These represent an array of values of the same type. They have a name, shape, and a datatype associated with them eg, sea surface temperature, precipitation, air pressure etc.
- Coordinate Variables: These are variables with the same name as that of the dimensions. They define a physical coordinate corresponding to that dimension.
- Attributes: NetCDF attributes store information about the data.
When I started dealing with NC files, which are datasets containing n-dimensions, variables, and attributes, I could no longer rely on Pandas. Because when your data starts getting complex with N-dimensions, and you can no longer classify it into simple rows and columns, Pandas be like,

Pandas has supported N-dimensional analysis in the past, in the form of Panels. However, support for Panels has been deprecated since version 0.20.0.
This is when I came across Xarray. Pandas favour Xarray for handling N-dimensional data in its documentation. There are built-in methods to convert data structures into Xarray and pandas, back and forth.
Xarray which is built upon pandas and NumPy provides two main data structures:
- DataArrays that wrap underlying data containers (e.g. NumPy arrays) and contain associated metadata.
- DataSets that are dictionary-like containers of DataArrays. It is very similar to the pandas’ data frame.
Time to play with some real data now!
To get started you’ll first have to install the prerequisites. Here’s a list:
- Python (3.8 or later)
- numpy (1.20 or later)
- pandas (1.3 or later)
- netcdf4 (Xarray needs this engine to deal with NetCDF files)
Importing a NetCDF file
To import data from a NetCDF file, use the open_dataset() method. You can also import multiple files at once in a single dataset using the open_mfdataset().
Let’s simply print the data set to see what that looks like –
import xarray as xr
try:
with xr.open_dataset('./temperature.nc') as ds:
print(ds)
except Exception as err:
print('oops...', err)
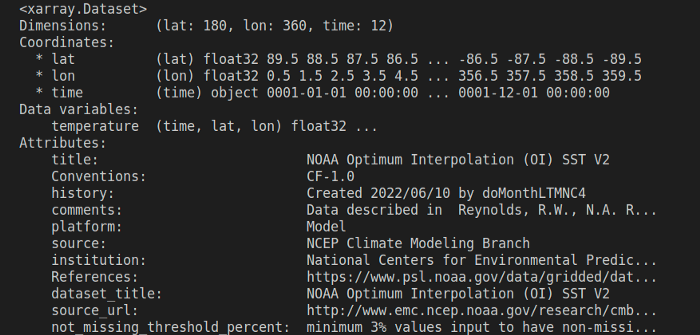
You can extract data from a particular variable simply using the dot operator. ds.<data_array_name>
ds.lat
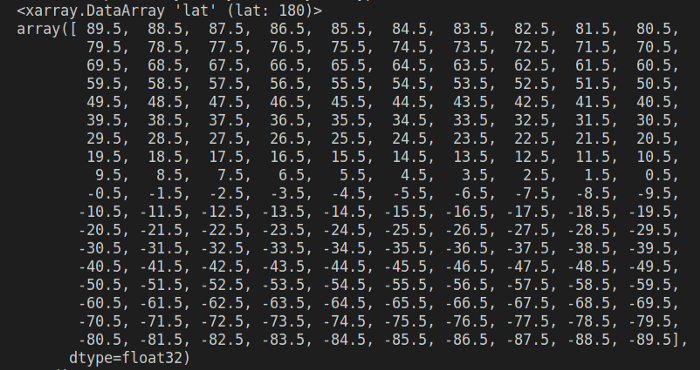
You can also query the dataset, using where()
ds.where(ds.temperature < -1)
E.g. To convert any Xarray dataset to a Pandas DataFrame, you can use
ds.to_dataframe()
Once you have a DataFrame you can apply any methods from pandas on it to get different views on the data
Dealing with Multiple datasets
Here’s how you can open multiple datasets at once and convert them to a DataFrame.
files_to_collate = ['temperature.nc', 'humidity.nc']
filters = 'temperature <= 0 & humidity > 50'
with xr.open_mfdataset(files_to_collate) as ds:
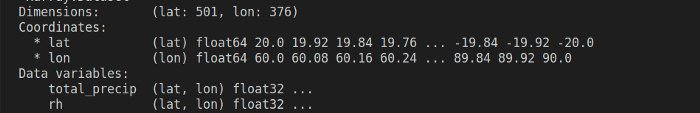
df = ds.to_dataframe().dropna(how="all")
filtered_df = df[df.eval(filters)]
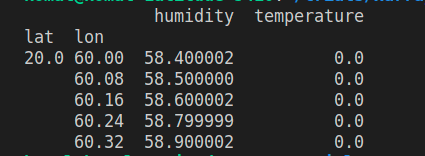
print(filtered_df)
The eval() function evaluates a string describing operations on Pandas DataFrame columns. The resulting DataFrame has columns from both the dataset variables, mapped against the coordinates variables. Smooth right?
There are many more methods to play around with Xarray datasets and data arrays.
This was a quick introduction to Xarray, readers are encouraged to explore more of these utilities as an exercise from the Xarray documentation.
Further, you can also plot the data arrays using Matplotlib.
If you’re interested in NetCDF data, You can download sample NC files from https://downloads.psl.noaa.gov/Datasets/noaa.oisst.v2/



